BiopathFx™
A Revolutionary New Way to Understand your DNA
In 2002, the first successful genome-wide association study (GWAS) was published showing a significant association between myocardial infarction and 5 variations in the genes for LTA, NFKBIL1, and BAT1. 1 These types of studies collect DNA from two groups of people – those with a disease or trait and those without – and subsequently scan millions of single-nucleotide polymorphisms (SNPs) across their genomes. Those SNPs that appear most frequently in the disease group are thought to be correlated with the pathology for that disease.
Genomic science gets sidetracked by GWAS
Fast forward to today: the scientific community was so emboldened by these first GWAS results that we now have nearly 4,000 new publications and 138,000+ disease / trait associations. Based on these results in combination with similar outcomes in gene-specific candidate-driven studies, savvy entrepreneurs the world over have created consumer genomics companies such as 23andMe, AncestryDNA, and MyHeritageDNA. These companies sample selected portions of the genome in micorarrays that target the SNPs observed in GWAS and similar studies, with the hopes of providing valuable, potentially actionable health-oriented information to consumers.
 To complement these microarrays, an even greater number of automated reports and “biohacking” tools have arisen to address the need for genetic interpretation, in many cases offering recommendations on how to improve one’s health – all in automated fashion.
To complement these microarrays, an even greater number of automated reports and “biohacking” tools have arisen to address the need for genetic interpretation, in many cases offering recommendations on how to improve one’s health – all in automated fashion.
The SNP-based nutrigenomics movement has become so widespread that its premise has been virtually unchallenged. Unfortunately, nucleotide polymorphisms are only partial contributors to an individual’s risk for developing a disease or unwanted health outcome. They cannot – in any meaningful way – be used, alone or in clusters, to deeply understand one’s genetic liabilities. The reason is simple…
Associating risks with population-specific changes in DNA, while proven to be statistically significant, does not capture the nuances and subtleties of genetic expression at the individual level.
 SNPs simply don’t tell you enough
SNPs simply don’t tell you enough
While you may share SNPs or groups of them with a particular population you belong to, your risk, at least according to GWA studies, is derived from only certain members of that population – and every single one of those people has a different genetic background. So while it may seem a valid assumption that you are at risk because portions of your DNA match other people that carry or demonstrate a disease or trait, it cannot possibly say enough about your own, unique circumstance. Why?
Because every genome is a snowflake – with its own set of liabilities and outcomes.
Depending on environment, lifestyle, and other external epigenetic pressures, those outcomes become even more distinct. Knowing this, it should be clear why SNPs, alone, cannot be used to derive conclusions about your health. In response to this challenge, we have aimed to create a revolutionary, new approach to interpreting DNA data, taking into consideration all genomic alterations that would change not only how human protein’s are transcribed, generated, and expressed – but also how they interact, distinctly within specific cells (i.e. tissues / organs) and pathways.
To this end, a novel process for individual genetic research and analysis was conceived….
 Introducing BiopathFx™
Introducing BiopathFx™
Our methodology embraces a vision for genetics-based “predictive medicine”. In order to realize this vision, several shortcomings in current interpretive methods needed to be addressed. The primary challenge faced when analyzing changes in an individual’s genome is the fact that genes are neither expressed in isolation nor uniformly throughout the body. Every person has their own, unique “phenotypic plasticity”, determining how each cell’s genetic expression will change in response to stimuli from the environment. This dynamic expression is further complicated by the interactions between the protein-products of genes in every tissue and organ. For this reason, we must not only know where changes have occurred in the germ-line (i.e. inherited genome), but also how those changes alter cellular processes in different ways, depending on their location or pathway.
After nearly a decade of research and experimentation, BiopathFx™ was created in 2015 to address these challenges. Instead of relying exclusively on population-based statistics to correlate risks with SNPs, we endeavored to take all genome-wide changes into consideration, including insertions, deletions, and SNPs – inside and outside coding regions. The result is a staggeringly accurate predicative model that will allow individuals to quickly and efficiently focus on the areas of their health that matter the most with maximum confidence.
How it works
Step 1
Your raw DNA data from a consumer DNA testing service of your choice is scanned into our system and insertions, deletions, and SNPs for over 20,000 protein-encoding human genes are extracted (depending on data source provided). For consumers with whole-genome sequences, additional areas are scanned including noncoding functional RNA, regulatory elements (e.g. transcription factors, enhancers, and promoters) and introns.2

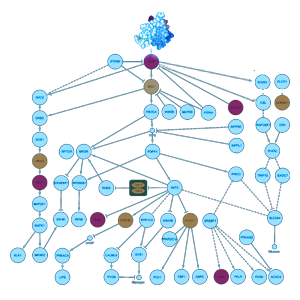
Step 2
Using a proprietary, cloud-based algorithm incorporating advanced predictive modeling, gene interactions are analyzed across hundreds of cellular, organismal, metabolic, and environmental pathways. Scores are derived reflecting potential impacts on health under specific endogenous and environmental influences / triggers.
Additional steps for those participating in the full program

Step 3
Lab results, comprehensive health history, and current and past symptoms are integrated to enrich all high-priority pathways. At this stage, we can make accurate predictions regarding the origins of your symptoms or health challenges.
Step 4
Millions of natural and pharmaceutical compounds are scanned for their epigenetic effects on each pathway’s genes. Instead of looking at gene-compound interactions in isolation as most other services do, we predict pathway-level effects, avoiding the pitfall of improving one problem while creating another – either in the same pathway or elsewhere in your biology.


Step 5
A comprehensive, individualized report is prepared that brings together your health history, lab results, symptoms, and goals into a cohesive whole, focusing on the pathways that are of highest importance for you. Each pathway description contains a breakdown of the genes used to derive its rating along with a detailed explanation of their potential effects on your health. Recommendations for lifestyle, diet, and supplementation are proposed, based on clinical studies, to help guide you to optimal health.
 What can BiopathFx™ do for you?
What can BiopathFx™ do for you?
By participating in our comprehensive technology-driven health optimization program, you will gain deep insight into your genetic template and grow in your understanding of how your genetic mutations play into both your physical and mental health.
Transcend Genomics employs neither licensed doctors nor medical specialists. Our services are for informational purposes only and are not intended to replace diagnosis or treatment by a qualified doctor, physician’s-assistant, nurse practitioner, chiropractor, nutritionist, or dietician. That being said, we also stand behind the value, depth, and breadth of scientific and medical data provided through our services and have received praise from people the world over for the impact it has made in their lives.
Though BiopathFx™ is still a new player on the genetics-based health scene, it has slowly but surely been gaining global recognition. All of the technologies employed in our methodology exist at various stages of maturity in the global bioinformatics community at large, but our application and customization of such technology is unique and even quite controversial. For this reason, it has not been formally positioned for use in clinical contexts, but instead is intended as a comprehensive tool to empower individuals to make well-informed decisions about their health.

